The proportion of progeny expected to be
 in the testcross of
in the testcross of
 , considering the given recombination frequencies and an interference value of 24%, is approximately 76.09%.
, considering the given recombination frequencies and an interference value of 24%, is approximately 76.09%.
To determine the proportion of
 we need to consider genetic linkage, recombination frequencies, and the concept of interference. Let's break it down step by step:
we need to consider genetic linkage, recombination frequencies, and the concept of interference. Let's break it down step by step:
1. Understanding the Cross:
- The initial cross is between two homozygous individuals with the genotype
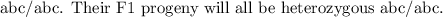
- A testcross involves crossing these F1 progeny with a homozygous recessive individual. However, since the problem doesn't specify the recessive alleles, we'll assume it's the same genotype
2. Genetic Map
- The map distance between genes a and b is 16 cM (centiMorgans), and between b and c is 9 cM.
3. Recombination Frequency:
- The recombination frequency between genes is approximately equal to the map distance (in cM) divided by 100. Therefore, the recombination frequency between a and b is 16%, and between b and c is 9%.
4. Interference and Coincidence:
- Interference is the phenomenon where a crossover in one region of a chromosome inhibits crossovers in an adjacent region. It's calculated as
 where coincidence is the observed double crossover rate divided by the expected double crossover rate.
where coincidence is the observed double crossover rate divided by the expected double crossover rate.
- Here, we have an interference of 24%, meaning the actual double crossover events are reduced by this amount.
5. Calculating Expected Double Crossovers:
- The expected double crossover frequency (without interference) between a and c is the product of the individual frequencies:
 or 1.44%.
or 1.44%.
- With 24% interference, the actual double crossover frequency is
6. Proportion of abc/abc Progeny:
- The progeny with the
 genotype result from either no crossover events or double crossovers between a and c .
genotype result from either no crossover events or double crossovers between a and c .
- The probability of no crossovers is simply the complement of any crossovers occurring, which is
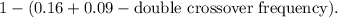
Let's calculate the exact proportion of
 progeny expected:
progeny expected:
The proportion of progeny expected to be
 in the testcross of
in the testcross of
 , considering the given recombination frequencies and an interference value of 24%, is approximately 76.09%. This accounts for both scenarios where no crossover events occur and scenarios involving double crossovers between genes a and c .
, considering the given recombination frequencies and an interference value of 24%, is approximately 76.09%. This accounts for both scenarios where no crossover events occur and scenarios involving double crossovers between genes a and c .